Overview of DVApred
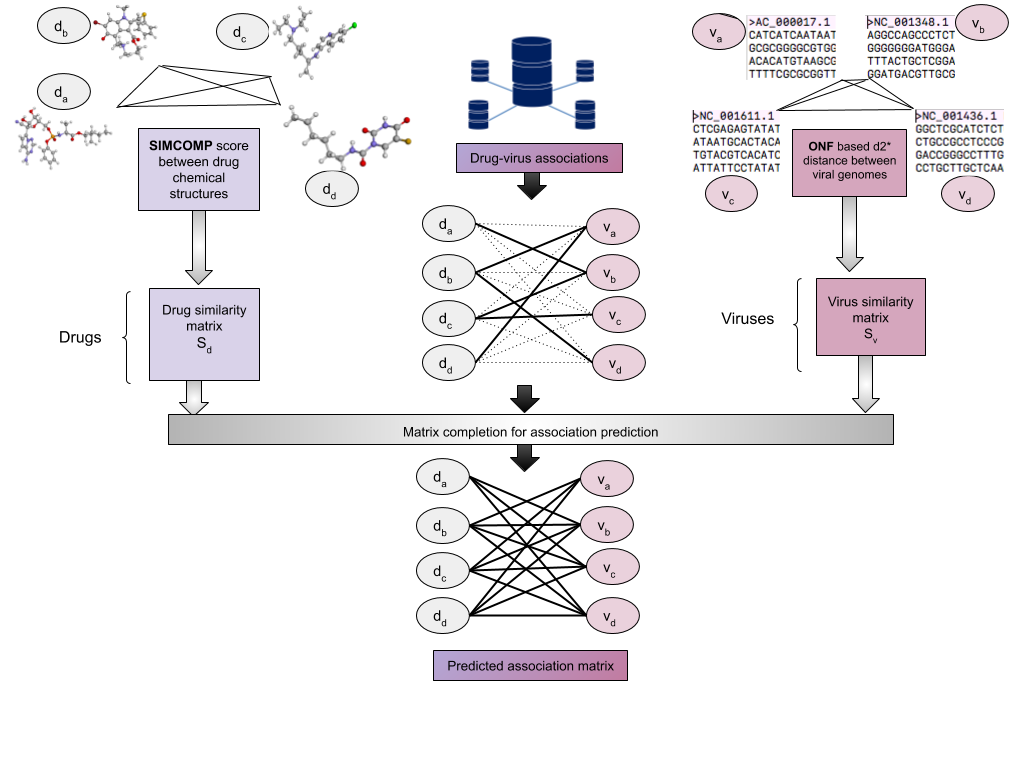
The above diagram depicts the schematic pipeline used in DVApred
The main contribution of this work is a manually curated database publicly shared, comprising of existing associations between viruses and their corresponding antivirals, obtained from DrugBank and other literature. The database gathers similarity information using the chemical structure of drugs by computaing the SIMCOMP score (using KEGG API) between them and the genomic sequence of viruses by finding d2* dissimilarity/distance (found using VirHostMatcher software). Along with this database, we make available a set of state-of-the-art computational drug re-positioning tools based on matrix completion.